
23 Sep Mount Sinai Icahn Researchers Develop Prediction Model for COVID-19 Mortality
MedicalResearch.com Interview with:

Dr. Pandey
Gaurav Pandey, Ph.D.
Assistant Professor
Department of Genetics and Genomic Sciences
Icahn Institute of Genomics and Multiscale Biology
Icahn School of Medicine at Mount Sinai, New York
MedicalResearch.com: What is the background for this study? What are the main findings?
Response: Given the toll that the COVID-19 pandemic has taken on people’s health and lives worldwide, it is crucial to be able to accurately predict patients’ outcomes, including their chances of mortality from the disease. Using the largest clinical dataset to date, and a systematical machine learning framework, the research team at Mount Sinai identified an accurate and parsimonious prediction model of COVID-19 mortality.
This model was based on only three routinely collected clinical features, namely patient’s age, minimum oxygen saturation over the course of their medical encounter, and type of patient encounter (inpatient vs outpatient and telehealth visits).
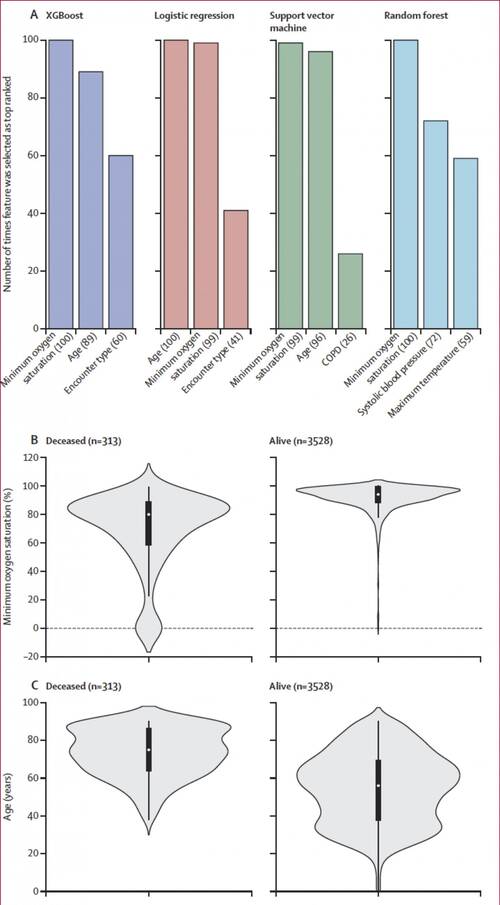
Top predictive features selected for the four classification algorithms (A) Top three predictive features identified using the recursive feature elimination method for the four classification algorithms across the 100 runs used to select the most discriminative features and train the corresponding candidate prediction models; the values in parentheses indicate the number of times the feature was selected as top ranked in the development dataset. Minimum oxygen saturation (B) and age (C) features, which were selected as top predictive features for all the four algorithms, are presented as violin plots showing the distributions of the values in the development dataset. In panels B and C, the black boxplots in the middle show the distribution of the values on the y axis, with the white dot indicating the median value; the width of the grey shape at a given value on the y axis indicates the probability of occurrence of that value in the population shown. The plots in panel B show that the median value (79%) of minimum oxygen saturation for the deceased group was significantly lower. CREDIT: Icahn School of Medicine at Mount Sinai
MedicalResearch.com: What should readers take away from your report?
Response: This model could yield an additional “vital sign” that is assessed regularly during a patient’s hospital course, that can be integrated into the clinical care flow of a COVID-19 patient. Clinical teams could use results from the prediction model throughout COVID-19 patients’ hospital courses to flag individuals at high risk of death so that they can promptly focus treatment and attention on such individuals to prevent their mortality.
MedicalResearch.com: What recommendations do you have for future research as a result of this work?
Response: In the future, it should be possible to develop more accurate prediction models for COVID-19 mortality and other outcomes by integrating multi-modal data collected from the patients. These data include demographics, co-morbidities, laboratory test measurements, vital signs, chest imaging, clinical notes and omic data, and can be integrated into prediction models using techniques like heterogeneous ensembles and deep learning.
This work was funded by NIH grants. None of the authors have any competing interests
Citations:
Clinical features of COVID-19 mortality: development and validation of a clinical prediction model
Arjun S Yadaw, PhDYan-chak Li, MPhilSonali Bose, MDProf Ravi Iyengar, PhD Prof Supinda Bunyavanich, MD Gaurav Pandey, PhD
October, 2020
DOI:https://doi.org/10.1016/S2589-7500(20)30217-X
[wysija_form id=”3″]
[last-modified]
The information on MedicalResearch.com is provided for educational purposes only, and is in no way intended to diagnose, cure, or treat any medical or other condition. Always seek the advice of your physician or other qualified health and ask your doctor any questions you may have regarding a medical condition. In addition to all other limitations and disclaimers in this agreement, service provider and its third party providers disclaim any liability or loss in connection with the content provided on this website.
Last Updated on September 23, 2020 by Marie Benz MD FAAD
