AHA Journals, Author Interviews, Heart Disease, Stroke / 12.03.2021
Stroke Risk: AI Can Help Predict Who Might Develop Atrial Fibrillation
MedicalResearch.com Interview with:
[caption id="attachment_56932" align="alignleft" width="200"] Dr. Fornwalt[/caption]
Brandon K Fornwalt, MD, PhD
Associate Professor, Director Department of Imaging Science and Innovation
Geisinger
MedicalResearch.com: What is the background for this study?
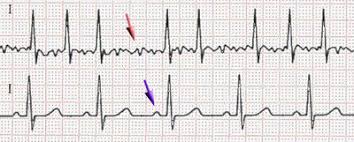
Response: Atrial fibrillation (AF) is an abnormal heart rhythm that is associated with outcomes such as stroke, heart failure and death. If we know a patient has atrial fibrillation, we can treat them to reduce the risk of stroke by nearly two-thirds. Unfortunately, patients often don’t know they have AF. They present initially with a stroke, and we have no chance to treat them before this happens. If we could predict who is at high risk of either currently having AF or developing it in the near future, we could intervene earlier and hopefully reduce bad outcomes like stroke. Artificial intelligence approaches may be able to help with this task.
Dr. Fornwalt[/caption]
Brandon K Fornwalt, MD, PhD
Associate Professor, Director Department of Imaging Science and Innovation
Geisinger
MedicalResearch.com: What is the background for this study?
Response: Atrial fibrillation (AF) is an abnormal heart rhythm that is associated with outcomes such as stroke, heart failure and death. If we know a patient has atrial fibrillation, we can treat them to reduce the risk of stroke by nearly two-thirds. Unfortunately, patients often don’t know they have AF. They present initially with a stroke, and we have no chance to treat them before this happens. If we could predict who is at high risk of either currently having AF or developing it in the near future, we could intervene earlier and hopefully reduce bad outcomes like stroke. Artificial intelligence approaches may be able to help with this task.
 Dr. Fornwalt[/caption]
Brandon K Fornwalt, MD, PhD
Associate Professor, Director Department of Imaging Science and Innovation
Geisinger
MedicalResearch.com: What is the background for this study?
Response: Atrial fibrillation (AF) is an abnormal heart rhythm that is associated with outcomes such as stroke, heart failure and death. If we know a patient has atrial fibrillation, we can treat them to reduce the risk of stroke by nearly two-thirds. Unfortunately, patients often don’t know they have AF. They present initially with a stroke, and we have no chance to treat them before this happens. If we could predict who is at high risk of either currently having AF or developing it in the near future, we could intervene earlier and hopefully reduce bad outcomes like stroke. Artificial intelligence approaches may be able to help with this task.
Dr. Fornwalt[/caption]
Brandon K Fornwalt, MD, PhD
Associate Professor, Director Department of Imaging Science and Innovation
Geisinger
MedicalResearch.com: What is the background for this study?
Response: Atrial fibrillation (AF) is an abnormal heart rhythm that is associated with outcomes such as stroke, heart failure and death. If we know a patient has atrial fibrillation, we can treat them to reduce the risk of stroke by nearly two-thirds. Unfortunately, patients often don’t know they have AF. They present initially with a stroke, and we have no chance to treat them before this happens. If we could predict who is at high risk of either currently having AF or developing it in the near future, we could intervene earlier and hopefully reduce bad outcomes like stroke. Artificial intelligence approaches may be able to help with this task.



 Dr. Helen Marsden PhD
Skin Analytics Limited
London, United Kingdom
MedicalResearch.com: What is the background for this study?
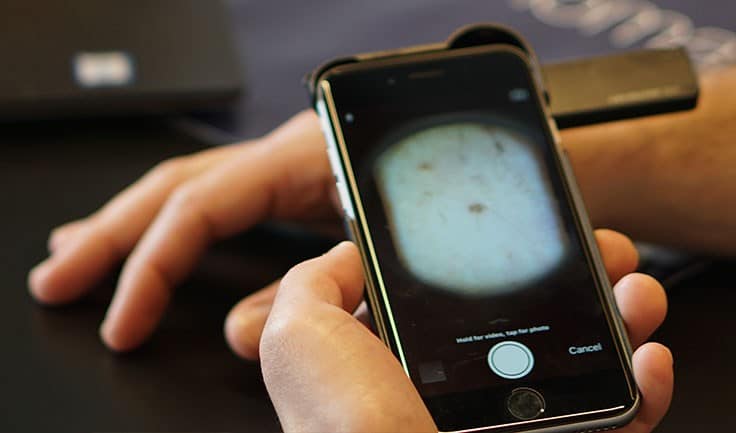
Response: In this technology age, with the explosion of interest and applications using Artificial Intelligence, it is easy to accept the output of a technology-based test - such as a smartphone app designed to identify skin cancer - without thinking too much about it. In reality, technology is only as good as the way it has been developed, tested and validated. In particular, AI algorithms are prone to a lack of “generalisation” - i.e. their performance drops when presented with data it has not seen before. In the medical field, and particularly in areas where AI is being developed to direct a patient’s diagnosis or care, this is particularly problematic. Inappropriate diagnosis or advice to patients can lead to false reassurance, heightened concern and pressure on NHS services, or worse. It is concerning, therefore, that there are a large number of smartphone apps available that provide an assessment of skin lesions, including some that provide an estimate of the probability of malignancy, that have not been assessed for diagnostic accuracy.
Skin Analytics has developed an AI-based algorithm, named: Deep Ensemble for Recognition of Malignancy (DERM), for use as a decision support tool for healthcare providers. DERM determines the likelihood of skin cancer from dermoscopic images of skin lesions. It was developed using deep learning techniques that identify and assess features of these lesions which are associated with melanoma, using over 7,000 archived dermoscopic images. Using these images, it was shown to identify melanoma with similar accuracy to specialist physicians. However, to prove the algorithm could be used in a real life clinical setting, Skin Analytics set out to conduct a clinical validation study.
Dr. Helen Marsden PhD
Skin Analytics Limited
London, United Kingdom
MedicalResearch.com: What is the background for this study?
Response: In this technology age, with the explosion of interest and applications using Artificial Intelligence, it is easy to accept the output of a technology-based test - such as a smartphone app designed to identify skin cancer - without thinking too much about it. In reality, technology is only as good as the way it has been developed, tested and validated. In particular, AI algorithms are prone to a lack of “generalisation” - i.e. their performance drops when presented with data it has not seen before. In the medical field, and particularly in areas where AI is being developed to direct a patient’s diagnosis or care, this is particularly problematic. Inappropriate diagnosis or advice to patients can lead to false reassurance, heightened concern and pressure on NHS services, or worse. It is concerning, therefore, that there are a large number of smartphone apps available that provide an assessment of skin lesions, including some that provide an estimate of the probability of malignancy, that have not been assessed for diagnostic accuracy.
Skin Analytics has developed an AI-based algorithm, named: Deep Ensemble for Recognition of Malignancy (DERM), for use as a decision support tool for healthcare providers. DERM determines the likelihood of skin cancer from dermoscopic images of skin lesions. It was developed using deep learning techniques that identify and assess features of these lesions which are associated with melanoma, using over 7,000 archived dermoscopic images. Using these images, it was shown to identify melanoma with similar accuracy to specialist physicians. However, to prove the algorithm could be used in a real life clinical setting, Skin Analytics set out to conduct a clinical validation study.











