Abuse and Neglect, Author Interviews, Gender Differences, Genetic Research, Science, University of Michigan / 05.01.2024
Bisexuality: Genetic Variants May Provide Reproductive Advantages
MedicalResearch.com Interview with:
[caption id="attachment_61209" align="alignleft" width="150"] Dr. Zhang[/caption]
Jianzhi "George" ZhangMarshall W. Nirenberg Collegiate ProfessorDepartment of Ecology and Evolutionary BiologyUniversity of MichiganAnn Arbor, MI 48109-1085
MedicalResearch.com: What is the background for this study?
Response: A few percent of humans perform same-sex sexual behavior (SSB), a trait that is partially heritable. Because SSB leads to fewer children, the stable maintenance of SSB-associated alleles in populations has been a long-standing Darwinian paradox.
A number of hypotheses have been proposed to resolve this paradox, but most of them lack clear empirical evidence. One version of the antagonistic pleiotropy hypothesis posits that SSB-associated alleles are subject to heterosexual advantage. Specifically, it was found that SSB-associated alleles are associated with more sexual partners when in heterosexuals (individuals of exclusive opposite-sex sexual behavior), which could lead to more offspring, potentially compensating the reduced reproduction of SSB individuals. While the above mechanism has likely worked in premodern societies, our recent study (PNAS 2023) found that it is no longer working in the modern United Kingdom, because the widespread use of contraception has decoupled the number of offspring from the number of sexual partners in heterosexuals.
Dr. Zhang[/caption]
Jianzhi "George" ZhangMarshall W. Nirenberg Collegiate ProfessorDepartment of Ecology and Evolutionary BiologyUniversity of MichiganAnn Arbor, MI 48109-1085
MedicalResearch.com: What is the background for this study?
Response: A few percent of humans perform same-sex sexual behavior (SSB), a trait that is partially heritable. Because SSB leads to fewer children, the stable maintenance of SSB-associated alleles in populations has been a long-standing Darwinian paradox.
A number of hypotheses have been proposed to resolve this paradox, but most of them lack clear empirical evidence. One version of the antagonistic pleiotropy hypothesis posits that SSB-associated alleles are subject to heterosexual advantage. Specifically, it was found that SSB-associated alleles are associated with more sexual partners when in heterosexuals (individuals of exclusive opposite-sex sexual behavior), which could lead to more offspring, potentially compensating the reduced reproduction of SSB individuals. While the above mechanism has likely worked in premodern societies, our recent study (PNAS 2023) found that it is no longer working in the modern United Kingdom, because the widespread use of contraception has decoupled the number of offspring from the number of sexual partners in heterosexuals.
 Dr. Zhang[/caption]
Jianzhi "George" ZhangMarshall W. Nirenberg Collegiate ProfessorDepartment of Ecology and Evolutionary BiologyUniversity of MichiganAnn Arbor, MI 48109-1085
MedicalResearch.com: What is the background for this study?
Response: A few percent of humans perform same-sex sexual behavior (SSB), a trait that is partially heritable. Because SSB leads to fewer children, the stable maintenance of SSB-associated alleles in populations has been a long-standing Darwinian paradox.
A number of hypotheses have been proposed to resolve this paradox, but most of them lack clear empirical evidence. One version of the antagonistic pleiotropy hypothesis posits that SSB-associated alleles are subject to heterosexual advantage. Specifically, it was found that SSB-associated alleles are associated with more sexual partners when in heterosexuals (individuals of exclusive opposite-sex sexual behavior), which could lead to more offspring, potentially compensating the reduced reproduction of SSB individuals. While the above mechanism has likely worked in premodern societies, our recent study (PNAS 2023) found that it is no longer working in the modern United Kingdom, because the widespread use of contraception has decoupled the number of offspring from the number of sexual partners in heterosexuals.
Dr. Zhang[/caption]
Jianzhi "George" ZhangMarshall W. Nirenberg Collegiate ProfessorDepartment of Ecology and Evolutionary BiologyUniversity of MichiganAnn Arbor, MI 48109-1085
MedicalResearch.com: What is the background for this study?
Response: A few percent of humans perform same-sex sexual behavior (SSB), a trait that is partially heritable. Because SSB leads to fewer children, the stable maintenance of SSB-associated alleles in populations has been a long-standing Darwinian paradox.
A number of hypotheses have been proposed to resolve this paradox, but most of them lack clear empirical evidence. One version of the antagonistic pleiotropy hypothesis posits that SSB-associated alleles are subject to heterosexual advantage. Specifically, it was found that SSB-associated alleles are associated with more sexual partners when in heterosexuals (individuals of exclusive opposite-sex sexual behavior), which could lead to more offspring, potentially compensating the reduced reproduction of SSB individuals. While the above mechanism has likely worked in premodern societies, our recent study (PNAS 2023) found that it is no longer working in the modern United Kingdom, because the widespread use of contraception has decoupled the number of offspring from the number of sexual partners in heterosexuals.


 Dr. Belloy[/caption]
Michael E. Belloy, PhD
Department of Neurology and Neurological Sciences
Stanford University, Stanford, California
MedicalResearch.com: What is the background for this study?
Response: Apolipoprotein E (APOE)*2 and APOE*4 are, respectively, the strongest protective and risk-increasing, genetic variants for late-onset Alzheimer disease. As such, one’s APOE genotype is highly relevant towards clinical trial design and Alzheimer’s disease research. However, most insights so far are focused on the associations of these APOE genotypes with Alzheimer’s disease risk in non-Hispanic white individuals.
One important aspect of our work is that we really increased sample sizes for non-Hispanic Black, Hispanic, and East Asian individuals, so that we now have better understanding of the associations of APOE genotypes with Alzheimer’s disease risk in these groups. In complement, we also did the largest investigation to date on the role of ancestry on the associations of APOE genotypes with Alzheimer’s disease risk. The scale of our study was thus a critical factor in generating novel insights.
Dr. Belloy[/caption]
Michael E. Belloy, PhD
Department of Neurology and Neurological Sciences
Stanford University, Stanford, California
MedicalResearch.com: What is the background for this study?
Response: Apolipoprotein E (APOE)*2 and APOE*4 are, respectively, the strongest protective and risk-increasing, genetic variants for late-onset Alzheimer disease. As such, one’s APOE genotype is highly relevant towards clinical trial design and Alzheimer’s disease research. However, most insights so far are focused on the associations of these APOE genotypes with Alzheimer’s disease risk in non-Hispanic white individuals.
One important aspect of our work is that we really increased sample sizes for non-Hispanic Black, Hispanic, and East Asian individuals, so that we now have better understanding of the associations of APOE genotypes with Alzheimer’s disease risk in these groups. In complement, we also did the largest investigation to date on the role of ancestry on the associations of APOE genotypes with Alzheimer’s disease risk. The scale of our study was thus a critical factor in generating novel insights.
 Dr. Stark[/caption]
Dr Mitchell Stark
Dr. Stark[/caption]
Dr Mitchell Stark
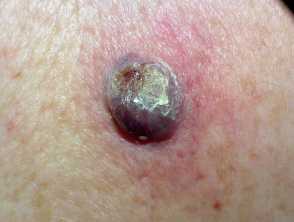
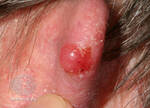 One example of a nodular melanoma without pigment.
One example of a nodular melanoma without pigment. 

 Dr. Davis[/caption]
Prof. Jonathan Davis, MD,
Chief of Newborn Medicine
Tufts Medical Center and
[caption id="attachment_60599" align="alignleft" width="125"]
Dr. Davis[/caption]
Prof. Jonathan Davis, MD,
Chief of Newborn Medicine
Tufts Medical Center and
[caption id="attachment_60599" align="alignleft" width="125"]

 Response: We were broadly interested in discovering instances of bacterial genes that have been acquired by diverse animal genomes over millions of years of evolution by the process of horizontal gene transfer (HGT). Since these events are quite rare and most previous discoveries have been serendipitous, we developed computational methods to identify genes acquired by HGT in animals. One of the exciting discoveries from our work was that vertebrate IRBP appeared to have originated in bacteria and is now a critical component of the vertebrate visual cycle, so this paper focuses on that one discovery.
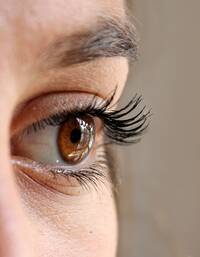
IRBP or interphotoreceptor retinoid binding protein is an important protein present in the space between two major cell types in our eyes, photoreceptor cells and RPE cells. Our ability to see involves an intricate set of steps where light is first sensed by causing a change (isomerization) in the chemical structure of molecules in the eye called retinoids. This sensing of light occurs in our photoreceptor cells. Following this change in the chemical structure, the retinoid needs to be recycled back to the chemical structure that can again sense light. This recycling occurs in RPE cells. IRBP performs the essential function of shuttling retinoids between the photoreceptors and the RPE cells, which allows the cycle of sensing and regeneration to work. Supporting its importance, mutations in IRBP (also known as retinol binding protein 3 or RBP3) can cause several severe human eye diseases.
Response: We were broadly interested in discovering instances of bacterial genes that have been acquired by diverse animal genomes over millions of years of evolution by the process of horizontal gene transfer (HGT). Since these events are quite rare and most previous discoveries have been serendipitous, we developed computational methods to identify genes acquired by HGT in animals. One of the exciting discoveries from our work was that vertebrate IRBP appeared to have originated in bacteria and is now a critical component of the vertebrate visual cycle, so this paper focuses on that one discovery.
IRBP or interphotoreceptor retinoid binding protein is an important protein present in the space between two major cell types in our eyes, photoreceptor cells and RPE cells. Our ability to see involves an intricate set of steps where light is first sensed by causing a change (isomerization) in the chemical structure of molecules in the eye called retinoids. This sensing of light occurs in our photoreceptor cells. Following this change in the chemical structure, the retinoid needs to be recycled back to the chemical structure that can again sense light. This recycling occurs in RPE cells. IRBP performs the essential function of shuttling retinoids between the photoreceptors and the RPE cells, which allows the cycle of sensing and regeneration to work. Supporting its importance, mutations in IRBP (also known as retinol binding protein 3 or RBP3) can cause several severe human eye diseases.

 Dr. Tauscher-Wisniewski,[/caption]
Sitra Tauscher-Wisniewski, MD
Vice President Clinical Development & Analytics
Novartis Gene Therapies
MedicalResearch.com: What is the background for this study? Would you briefly describe the condition of Spinal muscular atrophy (SMA)?
Response: At the 2023 Muscular Dystrophy Association Conference, we presented new data from two of our Long-Term Follow-Up (LTFU) studies, LT001 and LT002, which show the continued efficacy and durability of Zolgensma across a range of patient populations, with an overall benefit-risk profile that remains favorable. LT001 is a 15-year ongoing observational LTFU study following the Phase 1 START patients, who were the very first patients to receive our gene replacement therapy. LT-002 is a voluntary Phase 4 15-year ongoing follow-up safety and efficacy study of Zolgensma IV and investigational intrathecal (IT) OAV101 in patients previously treated in the Phase 3 IV studies (STR1VE-US, STR1VE-EU, STR1VE-AP, SPR1NT) and the Phase 1 IT study (STRONG).
Spinal muscular atrophy (SMA) is a rare, devastating genetic disease that leads to progressive muscle weakness, paralysis, and when left untreated in one of its most severe forms (SMA Type 1), permanent ventilation or death in 90% of cases by age 2. It is caused by a lack of a functional survival motor neuron 1 (SMN1) gene, and in the most severe forms results in the rapid and irreversible loss of motor neurons, affecting muscle functions, including breathing, swallowing and basic movement.
Dr. Tauscher-Wisniewski,[/caption]
Sitra Tauscher-Wisniewski, MD
Vice President Clinical Development & Analytics
Novartis Gene Therapies
MedicalResearch.com: What is the background for this study? Would you briefly describe the condition of Spinal muscular atrophy (SMA)?
Response: At the 2023 Muscular Dystrophy Association Conference, we presented new data from two of our Long-Term Follow-Up (LTFU) studies, LT001 and LT002, which show the continued efficacy and durability of Zolgensma across a range of patient populations, with an overall benefit-risk profile that remains favorable. LT001 is a 15-year ongoing observational LTFU study following the Phase 1 START patients, who were the very first patients to receive our gene replacement therapy. LT-002 is a voluntary Phase 4 15-year ongoing follow-up safety and efficacy study of Zolgensma IV and investigational intrathecal (IT) OAV101 in patients previously treated in the Phase 3 IV studies (STR1VE-US, STR1VE-EU, STR1VE-AP, SPR1NT) and the Phase 1 IT study (STRONG).
Spinal muscular atrophy (SMA) is a rare, devastating genetic disease that leads to progressive muscle weakness, paralysis, and when left untreated in one of its most severe forms (SMA Type 1), permanent ventilation or death in 90% of cases by age 2. It is caused by a lack of a functional survival motor neuron 1 (SMN1) gene, and in the most severe forms results in the rapid and irreversible loss of motor neurons, affecting muscle functions, including breathing, swallowing and basic movement.


 Dr. Levin[/caption]
Trevor Levin Ph.D.
Founder and CEO of Convergent Genomics that produces the Uroamp assay
San Francisco, CA
MedicalResearch.com: What is the background for this study?
Response: Bladder cancer is one of the most expensive and challenging to diagnose and treat. Therefore, identifying cost-effective urine bladder cancer biomarkers to complement or replace the gold-standard invasive and costly cystoscopy for the early detection and monitoring of this highly recurrent disease is crucial. At the international Agency for research on Cancer (IARC-WHO), we have developed a simple urine-based assay TERT promoter mutations, the most common mutations in bladder cancer, and showed that the urine biomarker could detect bladder cancer patients at diagnosis but many years prior to clinical diagnosis. However, in this study, we wanted to see whether a more comprehensive genomic profiling of urine samples collected years prior to clinical diagnosis of bladder cancer could identify even more patients before they develop any symptoms.
The study was based on the UroAmp test, a general urine test that identifies mutations in 60 genes, developed by the Oregon Health Science University spin out company, Convergent Genomics. Drawing on previous research to identify genetic mutations linked to bladder cancer, the research team narrowed the new test down to focus on mutations within just ten genes.
Working with colleagues from the Tehran University of Medical Sciences in Iran, they trialled the potential new test using samples from the Golestan Cohort Study, which has tracked the health of more than 50,000 participants over ten years, all of whom provided urine samples at recruitment. Forty people within the study developed bladder cancer during that decade, and the team were able to test urine samples from twenty-nine of them, along with samples from 98 other similar participants as controls.
Dr. Levin[/caption]
Trevor Levin Ph.D.
Founder and CEO of Convergent Genomics that produces the Uroamp assay
San Francisco, CA
MedicalResearch.com: What is the background for this study?
Response: Bladder cancer is one of the most expensive and challenging to diagnose and treat. Therefore, identifying cost-effective urine bladder cancer biomarkers to complement or replace the gold-standard invasive and costly cystoscopy for the early detection and monitoring of this highly recurrent disease is crucial. At the international Agency for research on Cancer (IARC-WHO), we have developed a simple urine-based assay TERT promoter mutations, the most common mutations in bladder cancer, and showed that the urine biomarker could detect bladder cancer patients at diagnosis but many years prior to clinical diagnosis. However, in this study, we wanted to see whether a more comprehensive genomic profiling of urine samples collected years prior to clinical diagnosis of bladder cancer could identify even more patients before they develop any symptoms.
The study was based on the UroAmp test, a general urine test that identifies mutations in 60 genes, developed by the Oregon Health Science University spin out company, Convergent Genomics. Drawing on previous research to identify genetic mutations linked to bladder cancer, the research team narrowed the new test down to focus on mutations within just ten genes.
Working with colleagues from the Tehran University of Medical Sciences in Iran, they trialled the potential new test using samples from the Golestan Cohort Study, which has tracked the health of more than 50,000 participants over ten years, all of whom provided urine samples at recruitment. Forty people within the study developed bladder cancer during that decade, and the team were able to test urine samples from twenty-nine of them, along with samples from 98 other similar participants as controls.


 Dr. Mosley[/caption]
Jonathan Mosley, MD, PhD
Associate Professor
Division of Clinical Pharmacology
Departments of Internal Medicine and Biomedical Informatics
Vanderbilt University Medical Center
MedicalResearch.com: What is the background for this study?
Response: Prostate cancer is an important source of morbidity and mortality among men. Earlier detection of disease is essential to reduce these adverse outcomes. Prostate cancer is heritable, and many single nucleotide polymorphisms (SNPs) associated with disease risk have been identified. Thus, there is considerable interest in using tools such as polygenic risk scores, which measure the burden of genetic risk variants an individual carries, to identify men at elevated risk of disease.
Dr. Mosley[/caption]
Jonathan Mosley, MD, PhD
Associate Professor
Division of Clinical Pharmacology
Departments of Internal Medicine and Biomedical Informatics
Vanderbilt University Medical Center
MedicalResearch.com: What is the background for this study?
Response: Prostate cancer is an important source of morbidity and mortality among men. Earlier detection of disease is essential to reduce these adverse outcomes. Prostate cancer is heritable, and many single nucleotide polymorphisms (SNPs) associated with disease risk have been identified. Thus, there is considerable interest in using tools such as polygenic risk scores, which measure the burden of genetic risk variants an individual carries, to identify men at elevated risk of disease.



 Dr. Mahdavi[/caption]
Dr. Sara Mahdavi, PhD
Clinical Scientist and Clinical Instructor
Research Appointment in the Faculty of Medicine
University of Toronto
Toronto, ON
MedicalResearch.com: What is the background for this study?
Response: This was a long-term study spanning 16 years and began with a population of young adults who were medically assessed on a regular basis. It was remarkable to see just how striking the effects of coffee were in the group that had the susceptible genetic variant, what we termed “slow caffeine metabolizers” yet no effect whatsoever in those who did not were termed “fast metabolizers”.
Dr. Mahdavi[/caption]
Dr. Sara Mahdavi, PhD
Clinical Scientist and Clinical Instructor
Research Appointment in the Faculty of Medicine
University of Toronto
Toronto, ON
MedicalResearch.com: What is the background for this study?
Response: This was a long-term study spanning 16 years and began with a population of young adults who were medically assessed on a regular basis. It was remarkable to see just how striking the effects of coffee were in the group that had the susceptible genetic variant, what we termed “slow caffeine metabolizers” yet no effect whatsoever in those who did not were termed “fast metabolizers”.




 Dr. den Hoed[/caption]
Marcel den Hoed, PhD
Researcher,Department of Immunology, Genetics and Pathology
Uppsala University
MedicalResearch.com: What is the background for this study? What are the main findings?
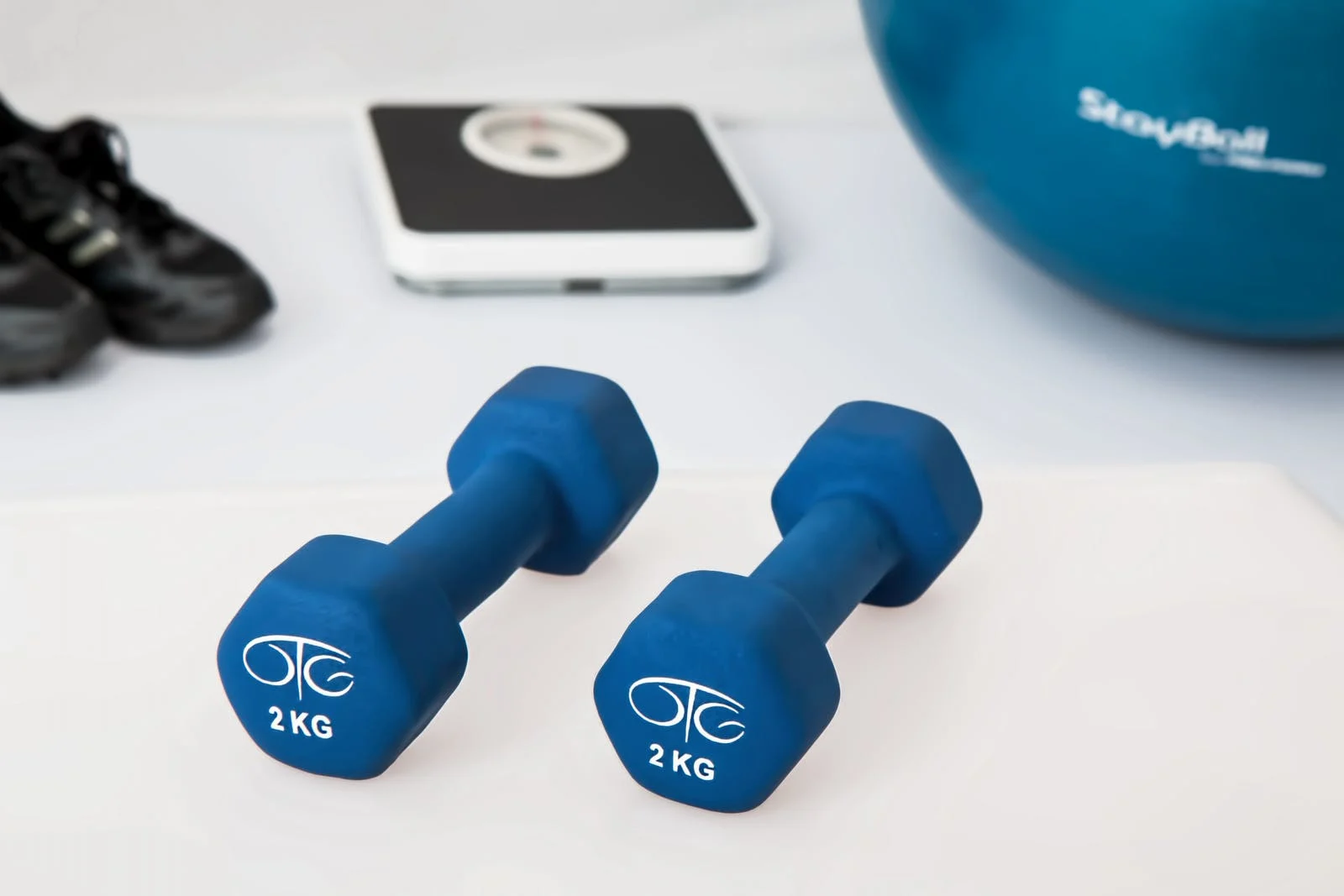
Response: In this paper we performed a multi-ancestry meta-analysis of 51 genome-wide association studies, in data from over 700,000 individuals. This yielded 11 DNA regions that are robustly associated with self-reported moderate-to-vigorous intensity physical activity during leisure time (MVPA), and 88 DNA regions for self-reported leisure screen time (LST).
Around half of the identified DNA regions are also associated with objectively assessed physical activity traits in data from the UK Biobank. Causal inference using a Mendelian randomization approach subsequently showed bidirectional causal effects between LST and body mass index (BMI), with the effect of LST on BMI being 2-3-fold larger than vice versa. Less LST and more MVPA protect from diabetes, attention deficit hyperactivity disorder, depression, and earlier age at death, with all causal effects of MVPA and leisure screen time being mediated or confounded by BMI. Further analyses showed that DNA regions associated with LST are more often located close to genes whose expression in skeletal muscle is altered by strength training than expected by chance, suggesting that these genes may influence the likelihood of adopting an active lifestyle by influencing the response to training.
Dr. den Hoed[/caption]
Marcel den Hoed, PhD
Researcher,Department of Immunology, Genetics and Pathology
Uppsala University
MedicalResearch.com: What is the background for this study? What are the main findings?
Response: In this paper we performed a multi-ancestry meta-analysis of 51 genome-wide association studies, in data from over 700,000 individuals. This yielded 11 DNA regions that are robustly associated with self-reported moderate-to-vigorous intensity physical activity during leisure time (MVPA), and 88 DNA regions for self-reported leisure screen time (LST).
Around half of the identified DNA regions are also associated with objectively assessed physical activity traits in data from the UK Biobank. Causal inference using a Mendelian randomization approach subsequently showed bidirectional causal effects between LST and body mass index (BMI), with the effect of LST on BMI being 2-3-fold larger than vice versa. Less LST and more MVPA protect from diabetes, attention deficit hyperactivity disorder, depression, and earlier age at death, with all causal effects of MVPA and leisure screen time being mediated or confounded by BMI. Further analyses showed that DNA regions associated with LST are more often located close to genes whose expression in skeletal muscle is altered by strength training than expected by chance, suggesting that these genes may influence the likelihood of adopting an active lifestyle by influencing the response to training.





 Dr. Ying-Hui Fu[/caption]
Ying-Hui Fu, PhD
Professor, Neurology
Weill Institute for Neurosciences
UCSF
MedicalResearch.com: What is the background for this study?
Response: Most people are aware that a lack of sleep is associated with all sorts of health issues. However, familial natural short sleeper (FNSS) individuals sleep 4-6.5 hours a night most of their live and stay healthy. We set out to determine whether natural short sleep mutations can offer protection from various diseases. We chose Alzheimer as an example to start.
Dr. Ying-Hui Fu[/caption]
Ying-Hui Fu, PhD
Professor, Neurology
Weill Institute for Neurosciences
UCSF
MedicalResearch.com: What is the background for this study?
Response: Most people are aware that a lack of sleep is associated with all sorts of health issues. However, familial natural short sleeper (FNSS) individuals sleep 4-6.5 hours a night most of their live and stay healthy. We set out to determine whether natural short sleep mutations can offer protection from various diseases. We chose Alzheimer as an example to start.

 Dr. Torkamani[/caption]
Ali Torkamani, Ph.D.
Director of Genomics and Genome Informatics
Scripps Research Translational Institute
Professor, Integrative Structural and Computational Biology
Scripps Research
La Jolla, CA 92037
MedicalResearch.com: What is the background for this study?
Response: Prior research has shown that people with higher polygenic risk for coronary artery disease achieve greater risk reduction with statin or other lipid lowering therapy. In general, adherence to standard guidelines for lipid lowering therapy is low - about 30% of people who should be on lipid lowering therapy are, with no correlation to their genetic risk. We set out to see whether communicating personalized risk, including polygenic risk, for coronary artery disease would drive the adoption of lipid lowering therapy.
Dr. Torkamani[/caption]
Ali Torkamani, Ph.D.
Director of Genomics and Genome Informatics
Scripps Research Translational Institute
Professor, Integrative Structural and Computational Biology
Scripps Research
La Jolla, CA 92037
MedicalResearch.com: What is the background for this study?
Response: Prior research has shown that people with higher polygenic risk for coronary artery disease achieve greater risk reduction with statin or other lipid lowering therapy. In general, adherence to standard guidelines for lipid lowering therapy is low - about 30% of people who should be on lipid lowering therapy are, with no correlation to their genetic risk. We set out to see whether communicating personalized risk, including polygenic risk, for coronary artery disease would drive the adoption of lipid lowering therapy.
 Dr. Mapara[/caption]
Markus Y Mapara, MD
Professor of Medicine
Director of the Blood and Marrow Transplantation
Columbia University Medical Center
MedicalResearch.com: What is the background for this study? What are the main findings?
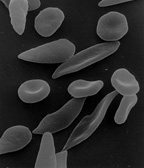
Response: Sickle cell disease is caused by a point mutation in the beta-globin gene of hemoglobin resulting in the production of abnormal hemoglobin which leads to formation of sickle-shaped RBC under conditions of low oxygen. Sickle cell disease affects about 100,000 patients in the US which are predominantly African American. The only curative approach is to perform an allogeneic bone marrow transplant which is however fraught with significant treatment-related risks if a matched sibling donor is not available.
The current study describes the successful application of a novel gene therapy to treat patients with sickle cell disease. The strategy is based on a gene-addition approach to introduce the genetic information for a Hemoglobin F-like molecule termed HgAT87Q into hematopoietic stem cells. The expression of this novel hemoglobin prevents polymerization of HgbS and has now been demonstrated to prevent the occurrence of vaso-occlusive pain crises in sickle cell disease patients.
Dr. Mapara[/caption]
Markus Y Mapara, MD
Professor of Medicine
Director of the Blood and Marrow Transplantation
Columbia University Medical Center
MedicalResearch.com: What is the background for this study? What are the main findings?
Response: Sickle cell disease is caused by a point mutation in the beta-globin gene of hemoglobin resulting in the production of abnormal hemoglobin which leads to formation of sickle-shaped RBC under conditions of low oxygen. Sickle cell disease affects about 100,000 patients in the US which are predominantly African American. The only curative approach is to perform an allogeneic bone marrow transplant which is however fraught with significant treatment-related risks if a matched sibling donor is not available.
The current study describes the successful application of a novel gene therapy to treat patients with sickle cell disease. The strategy is based on a gene-addition approach to introduce the genetic information for a Hemoglobin F-like molecule termed HgAT87Q into hematopoietic stem cells. The expression of this novel hemoglobin prevents polymerization of HgbS and has now been demonstrated to prevent the occurrence of vaso-occlusive pain crises in sickle cell disease patients.


